BACKGROUND: Several genetic markers have been associated with MS susceptibility; however, uncovering the genetic aetiology of the complex phenotypic expression of MS has been more difficult so far. The most common approach in imaging genetics is based on mass-univariate linear modelling (MULM), which faces several limitations.
“Wow, what the hell is MULM? Let’s find out!”
OBJECTIVE: This study applied a novel multivariate statistical model (more gobbledygook), sparse reduced-rank regression (sRRR), to identify possible associations of glutamate related single nucleotide polymorphisms (SNPs) (genetic differences) and multiple MRI-derived phenotypes in MS.
“A phenotype is a term we use to describe how a disease looks, for example clinical phenotypes are RRMS vs. SPMS vs. PPMS.”
METHODS: Seven phenotypes related to brain and lesion volumes for a total number of 326 relapsing-remitting and secondary-progressive MSeers and a total of 3809 glutamate related and control SNPs were analysed with sRRR, which resulted in a ranking of SNPs in decreasing order of importance (‘selection probability’). Lasso regression (statistical gobbledygook) and MULM were used as comparative statistical techniques to assess consistency of the most important associations over different statistical models.
RESULTS: Five SNPs within the NMDA-receptor-2A-subunit (GRIN2A) domain were identified by sRRR in association with normalized brain volume (NBV), normalized grey matter volume and normalized white matter volume (NMWM). The association between GRIN2A and both NBV and NWMV was confirmed in MULM and Lasso analysis.
CONCLUSIONS: Using a novel, multivariate regression model confirmed by two other statistical approaches we show associations between GRIN2A SNPs and phenotypic variation in NBV and NWMV in this first exploratory study. Replications in independent datasets are now necessary to validate these findings.
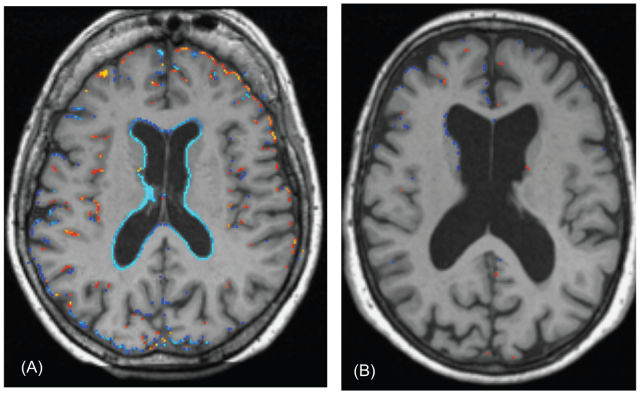 |
| Brain atrophy or shrinakge of the brain may be linked to the genetic control of glutamate levels. |
“In short this study is suggesting that genetic variants that control the amino acid glutamate levels in the brain are associated with brain volume (shrinkage) and white matter volume. Am I surprised? No, glutamate is an excitatory chemical and has been shown to speed up neurodegeneration in a lot of conditions; possibly MS.”
“Studies of this nature need to be reproduced and there is a high chance that these finding will not be reproduced. This is the nature of genetic studies describing associations; most findings are false positive findings. These investigators should really have reproduced their findings before publishing. So watch this this space. Any bets?”
How does brain atrophy typically progress in MS patients? Don't we all start losing volume early on, and then the loss accelerates or becomes more noticeable later in the disease course? Or do some of us get to keep more brain than others?
If I am right glutamate also stimulates the immunesystem.
I had a test on a bioimpedence machine not sure how scientific that is and it showed my dopamine off the scale but the tester said it was probarbly glutamate
Anon 10.53Glutamate can do many things including influence immune cells which have glutamate receptors such as metaboltrophic glutatamte receptors